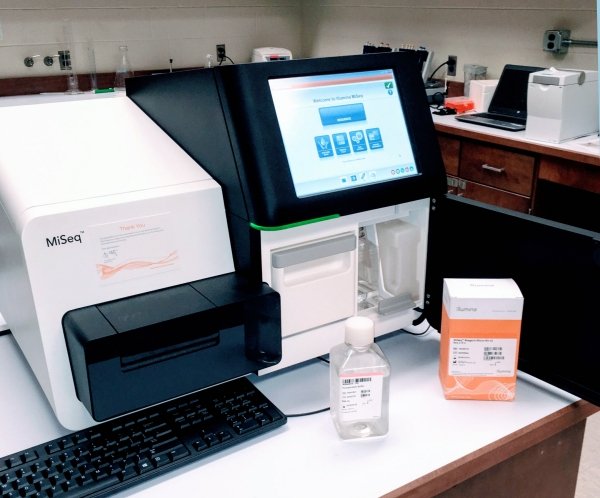
ODU offers DNA sequencing within our MiSeq Core Facility.
The Old Dominion University Sequencing Cost Center offers next generation sequencing (NGS) via the Illumina MiSeq™ desktop sequencing platform. This sequencer permits high throughput sequencing for a variety of genomic samples. Compatible kits allow targeted resequencing, small-genome sequencing, RNA sequencing, etc. with read lengths from 2x25 up to 2x300 and single-end reads from 1-25 Million (paired end 2-50M). Sequencing takes place one base at a time, by fluorescent detection of terminators at each step. Total run times depend upon the kit used and vary from less than a day to up to 3 days.
Sequencing clients purchase their own consumables (Library preparation kits, MiSeq kits, PhiX, BioAnalyzer chips, etc). Cost for services can be found below. To order services, please fill out the form. You will then be contacted by the technician for more details and sequence scheduling.
Recommendations
- Research the best type of library prep kit for your particular samples. Illumina tech services are very knowledgeable about this and have always been quite helpful. I recommend contacting them if you are not sure what to use.
- Similarly, if NGS is new to you, you should consult Illumina tech support when choosing the right MiSeq compatible kit for the actual sequencing. This will depend on your quantity of libraries, sample length, number of reads desired, etc.
- Appropriate and quality library preparation is critical. It is well worth it to check your samples on a bioanalyzer for quality and size and on a Qubit to quantify. Accurate sample normalization is necessary for a successful sequencing run.
- Prior to pooling your indexed libraries be certain they are diluted to equimolar concentrations. Pool libraries at a minimum of 10nM and be sure to keep at least half of this 10nM pooled library in your freezer. This will be important if you wish to rerun your samples for any reason.
- All runs require a spike of PhiX. This is a positive control that will allow us to determine if the run occurred properly. PhiX improves cluster generation and helps with sequencing and alignment. It is especially necessary when working with low-diversity libraries. PhiX can be the difference between a successful and a failed run.
It is highly recommended that MiSeq clients avail themselves of the support provided by Illumina for determining which products best suit the samples and research objectives of each individual client. There are an impressive variety of products available to address varied interests, therefore understanding your need and choosing the right options are important and simplified with these guides, selectors and other links:
MiSeq Sequencer Cost Rate Sheet
| User Type | MiSeq Run Fee | Technician Time |
|---|---|---|
| Internal | $50 per run | $30 per hour |
| External | $100 per run | $60 per hour |
Run Fees
Run fees cover the cost of maintenance and service for the instrument. Users will cover the costs of their own sequencing consumables (i.e., MiSeq Sequencing Kits, PhiX, BioAnalyzer chips, KAPA Quantification kits).
Technician time
Technician time charges will be quoted on a per-request basis.
Technician time associated with run set up, loading, and data handling is anticipated at approximately 2.5 hours ($75). Quality control analyses can be performed by the MiSeq Sequencer facility, the technician time associated with the BioAnalyzer is 2 hours ($60) and KAPA Quantification is 2.5 hours ($75).